Introduction to the Strawberry Clean Plant Program

For over 50 years, the Department of Plant Sciences Pomology Program at the Davis campus of the University of California (UC) has been the home of one of the most successful strawberry breeding programs in the world. Cultivars developed by the Plant Sciences Department are patented and licensed by UC Davis Technology Transfer Services, and maintained at and distributed by Foundation Plant Services (FPS).
Maintenance and distribution involves annual disease and identity testing, production and distribution of tissue culture meristem tip explants, and distribution of conventionally propagated plants produced off-site by special agreement. FPS tests and maintains advanced selections produced by UC strawberry breeders, ensuring the smooth release of disease-tested propagating stock when new cultivars are introduced.
Production of Meristem Plants

A 'meristem plant' is the term used in industry to refer to a plant grown in tissue culture from a meristem tip. The main reasons for producing meristem plants are to eliminate pathogens from planting material and to reinvigorate plants. Meristem plants produce many more daughter plants than are produced by conventionally-propagated plants. A ‘meristem tip’ consists of meristem tissue plus one or two leaf primordia, and is less than 0.5 mm in size. Meristem tissue is found in young growing points where cells are dividing but have not yet differentiated into leaves, stems or other organs. The following steps describe the process of producing strawberry ‘meristem plants.’ The process from mother plant to meristem plant in potting soil takes approximately one year.
1. Mother plants
The original strawberry plant material for a UC cultivar is provided by the UC strawberry breeder before release, and is tested repeatedly over several years before becoming a mother plant (also known as nuclear stock). Mother plants are propagated conventionally by runner propagation throughout the year.
2. Annual Disease Testing
To maintain their certification status, mother plants of each cultivar are inspected and tested annually for viruses using strawberry biological indicators and herbaceous hosts, in accordance with the California Department of Food & Agriculture’s (CDFA) Registration & Certification Program. They are also tested for viruses and other pathogens by polymerase chain reaction (PCR).
3. DNA Identification
The cultivar identity of each mother plant from which meristem tips are excised is verified using DNA ‘fingerprinting’ marker technology for quality control. Leaves are sampled prior to heat treatment.
4. Heat Treatment
Mother plants with attached daughter plants are placed into a heat treatment chamber for a minimum of 3 weeks at a constant 37C, 70% relative humidity and a 16-hour daylength. A strong plant with well-established roots is required to survive this treatment.
5. Tissue Culture
Daughter plants are harvested and meristem tips are excised from terminal and axillary buds on the crowns. Surface sterilization is not necessary if care is taken when removing the outer leaves. Each meristem tip is excised aseptically, in a laminar airflow transfer hood using a scalpel and forceps under an 80-100x zoom stereoscope. The scalpel is flame sterilized frequently. Meristem shoot tips are gently placed on MSI medium (half-strength Murashige and Skoog salts and vitamins, 2 % sucrose, 1mg/l IAA, 0.6% agar pH 5.8) on which they will usually form shoots and roots.
After roots form and the explant is approximately one cm. tall, the meristem plant is transferred to a maintenance medium without growth hormones (Murashige and Skoog salts and vitamins, 3% sucrose, 0.2% Phytagel, pH 5.8). The plants are transferred to fresh medium every four to eight weeks.
Plants destined for out-of-state or foreign countries are usually shipped in vitro (in tubes). Nurseries have the option of buying plants in vitro or established in soil if they can arrange to pick them up at FPS. Plants can also be stored in MS in tubes wrapped with Parafilm at 4° C for one to two years.
On MSI medium, we find that plants produce no or very little callus tissue. These strategies minimize the risk of generating an off-type plant. Similar conclusions were reached on a similar medium by Sobczykiewicz (1979). Occasionally, plants with leaf variegations or white-streak chimeras are observed in tubes or soil and are discarded.
The percent of meristem tips that successfully grow depends on cultivar and technician dexterity among other factors. Success averages 30 to 50%, although the range is as low as 10% for some cultivars such as ‘Seascape,’ and can reach 90 - 100% for other cultivars such as ‘Camarosa’ and ‘Albion.’
As daughter plants typically each have three to six meristem shoot tips, a mother plant that has 30 attached daughter plants can have more than 100 meristem shoot tips. The FPS program minimizes the possibility of producing a variant type plant by producing only a single meristem plant from a meristem shoot tip; each mother plant can therefore be expected to yield approximately 100 meristem plants.
Virus Elimination in Strawberries Videos
6. Acclimation
Plants for in-state orders are transferred from tubes to 2-inch pots with sterile potting mix and then gradually acclimated to ambient humidity in a humid chamber. The acclimation is critical and takes two to three weeks.
7. Greenhouse
Meristem plants are transplanted to 4-inch pots and moved to a greenhouse. They are inspected by the California Department of Food and Agriculture prior to being picked up by customers.
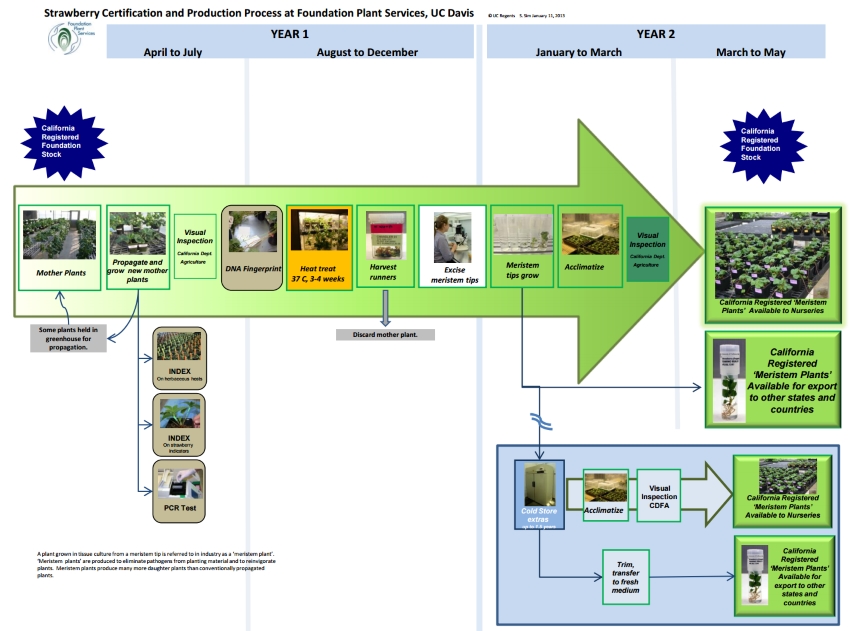
Click image for larger version
Strawberry Viruses
Strawberries are affected by over 30 viruses and phytoplasmas, many of which can greatly reduce yield, rapidly spread in the field, and may not cause obvious symptoms. Strawberry mottle virus (SMoV), Strawberry crinkle virus (SCV) and Strawberry mild yellow edge virus (SMYEV) are some of the most common viruses of strawberries.
Some severe strains of SMoV can reduce yield as much as 30%. Often, modern cultivars are symptomless when infected with just one virus but in a mixed infection with more than one virus, a decline disorder can develop in which the leaves turn red, and plants weaken and may die (Martin and Tzanetakis, 2006).
Symptoms of virus infection in susceptible indicator strawberry species can be very mild or severe, and include leaf distortion crinkling, and flecks or a mottled pattern. SMoV, SCV and SMYEV are aphid-transmitted; other viruses, including strawberry pallidosis associated virus (SPaV) and beet pseudo-yellows virus (BPYV), are whitefly-transmitted; and another group, including tomato ringspot virus (ToRSV), strawberry latent ringspot virus (SLRSV), and Arabis mosaic virus (ArMV), are nematode-transmitted.
One of the best ways to minimize loss due to virus infection is to plant healthy stock that has been certified by the California Department of Food & Agriculture or a similar government agency.
Why Tissue Culture is Used for Strawberry Clean Stock
 At FPS, tissue culture is used to eliminate viruses and other pathogens and to reinvigorate strawberry plants for increased runner production in nurseries. Multiplication of plants in tissue culture is avoided to minimize chances of variant type plants.
At FPS, tissue culture is used to eliminate viruses and other pathogens and to reinvigorate strawberry plants for increased runner production in nurseries. Multiplication of plants in tissue culture is avoided to minimize chances of variant type plants.
The use of tissue culture for the elimination of strawberry viruses was first reported in the 1960s and is used by certification programs worldwide. It is a common technique used successfully in many crop plants. Combined with heat treatment, tissue culture has been very successful in producing healthy strawberry planting stock. For many years at FPS, all of the plants produced from virus-positive mother plants have tested negative for virus after treatment.
Research and experience has shown that, even when viruses are not detected, a plant that has been through heat treatment and tissue culture is more vigorous and can produce many more daughter plants than conventionally propagated plants. One explanation is that this is because the plant regains juvenile traits which are thought to have a more vegetative rather than a fruiting growth habit. Runner productivity of tissue culture derived plants can be expected to be at least 50% more than conventionally propagated plants; reports range from no increase to 400% more (Swartz, et al. 1981; Scott et al. 1985; Merkle 1993).
One ‘meristem plant’ from the FPS program can bear between 300 to 800 daughter plants in one season in a screened house, depending on cultivar and growing conditions; up to 2000 daughter plants from one ‘meristem plant’ has been reported. This is in contrast to conventional runner propagation, in which each plant produces approximately 100 daughter plants in a season (C. Gaines, personal communication).
Multiplication of strawberry plants in tissue culture has mixed acceptance and is not used at FPS. Its advantages are reduced generation time and reduced exposure to insects and disease (Moisander et al. 2006). However, problems with variant types, especially a hyperflowering trait, have been documented (Jemmali 1995). These problems can be addressed by restricting the number of subcultures, reduced hormones in media, and fruiting trials of plants produced in tissue culture. (Martinelli 1992, Boxus et al. 2000).
Disease Testing
Each cultivar and advanced breeder selection is tested annually for over 15 pathogens (see list). Three standard methods are used to detect pathogens: graft indexing, herbaceous host indexing and polymerase chain reaction (PCR). These tests complement and serve as checks for each other. The graft and herbaceous host indexes are capable of screening large numbers of plants relatively inexpensively for the presence of a wide range of viruses. PCR is rapid, highly sensitive, and can identify which virus is present but requires expensive equipment and reagents, specific primers, and highly skilled technicians.
Graft index
Graft indexing is currently the only test required for the CDFA Registration and Certification Program. Each plant to be tested, referred to as the candidate plant, is grafted into three virus-sensitive strawberry clones used as indicator plants for strawberry viruses: Clone UC4, UC5 and UC10 or UC11. UC4 and UC5 are selections of hybrids of the wild strawberry Fragaria vesca. UC10 and UC11 are selections of the wild strawberry F. virginiana. The leaf blade of the candidate plant is trimmed and the petiole is cut into a wedge. The center leaflet on an indicator plant is removed and the petiole split to receive the leaflet from the candidate plant. Three grafts are placed on each indicator plant. The petiole is wrapped and the indicator plant is kept under mist to allow grafts to heal. If the candidate plant is virus infected, symptoms appear on the new growth of the indicator plant in approximately one month.
Herbaceous host index
Herbaceous host indexing is employed to test for the presence of sap-transmissible viruses. Four virus-sensitive herbaceous species Chenopodium amaranticolor, C. quinoa, Cucumis sativus and Nicotiana clevelandii are used. Young leaves of candidate plants are ground in buffer. Then, this plant extract is rubbed onto indicator plants pre-dusted with carborundum. This transmits virus particles that may be present in the candidate plant to the indicator plant. Indicator plants are incubated in a greenhouse. If the candidate plant is infected, symptoms appear in indicator plants in approximately 10-14 days.
Polymerase chain reaction (PCR) tests
Candidate plants are PCR tested for viruses, phytoplasmas and Xanthomonas fragariae. As more is known about the molecular biology of pathogens, and primers become available, tests are added to the program. Small tubes, each containing one plant sample and PCR test reagents for one pathogen, are put in a thermocycler for three hours. Reaction products are loaded into an agarose gel for gel electrophoresis lasting about one hour. The gel is stained with a UV-sensitive stain and photographed with UV light. If the candidate plant is infected, a bright band appears where stain binds to virus-specific nucleic acid. Each column represents one virus test for one plant sample. PCR test results can be obtained in one day.
Strawberry DNA Fingerprinting for Identity
New strawberry cultivars are regularly introduced to the strawberry industry, with no single cultivar dominating the market. More than 70 percent of the strawberries produced in California are from UC Davis cultivars. Strawberry growers rely on UC Davis Foundation Plant Services (FPS) to provide nurseries with virus-tested, correctly identified planting stock of the newest and best UC-patented strawberry cultivars. Historically, there has been no objective screening test to accurately distinguish cultivars. In response to industry and consumer needs, FPS scientists developed a DNA “fingerprinting” methodology to correctly identify strawberry cultivars.
In the fingerprinting process, DNA is extracted from fresh strawberry leaves, then specific marker DNA is amplified using polymerase chain reaction. The amplified marker DNA for each strawberry cultivar is unique, just like a human fingerprint. By analyzing the marker DNA and comparing the results to a newly developed database of strawberry DNA fingerprints, researchers can determine a strawberry plant’s varietal identity.
Simple Sequence Repeats (SSRs) are areas in the DNA of an organism in which the same small nucleotide sequence, usually consisting of two to six base pairs, are repeated many times. SSRs are abundant in many organisms, are widely used for genetic studies, and several, in combination, can distinguish closely related cultivars within a species. SSRs are also called microsatellites.
All mother plants that are used to produce meristem plants are DNA tested to confirm correct cultivar identification. An efficient and reliable DNA fingerprinting scheme using Simple Sequence Repeats (SSRs) was developed by FPS in cooperation with the USDA and made publicly available. Strawberry plant samples can now be sent to private labs to verify their identity. Nurseries are encouraged to check their stock regularly for varietal correctness.
Other biochemical and molecular marker systems have been used for strawberry cultivar identification, including isozymes (Bell and Simpson, 1994), Randomly Amplified Polymorphic DNA (RAPD) markers (Degani et al. 1998; Garcia et al. 2002; Hancock et al. 1994), Amplified Fragment Length Polymorphism (AFLP) markers (Miroslaw et al. 2002), and Inter Simple Sequence Repeat (ISSR) markers (Arnau et al. 2002). Each of these systems has technical limitations; all have known problems with reproducibility of data among labs.
Simple Sequence Repeats have been widely used for cultivar identification in grape, walnut and other clonally propagated crops (Dangl et al. 2001; Dangl et al. 2005), but SSR markers take considerable time and expertise to develop (Lewers et al. 2005). Application of this technology to strawberry is particularly difficult because strawberries are octaploid, with eight copies of each gene, and because inbreeding can lead to cultivars with multiple copies of the same allele at a single locus. Since SSR technology can detect only the presence or absence of an allele, but not copy number, there are often several genetic combinations that could lead to a particular profile.
This SSR marker system can distinguish between closely related UC-patented strawberry cultivars. Previously published markers were used to reduce development time; amplified fragments are scored as dominant markers. Because primer binding is highly specific, this system is both highly accurate and reproducible by other laboratories.
The basic procedure is to collect, dry and extract DNA from young leaves, although other tissues can also be used. The DNA is amplified using one optimal PCR condition. The PCR products are loaded into the genetic analyzer which results in a pattern of peaks that are scored as present or absent. The process is fast (about two days are required), accurate and very reproducible. The disadvantages are that it requires highly skilled technicians and expensive reagents and equipment.
Sixteen previously published primer pairs from strawberry were tested and described in A New System for Strawberry Cultivar Identification Developed at Foundation Plant Services (FPS). (Dangl, G.S. et al. 2007) This and other publications are available at http://fps.ucdavis.edu. Six primer pairs amplified consistently as well as produced variable peak sizes over our study set of 45 commonly grown strawberry cultivars and breeder’s selections. One primer pair, ARSFL-09, had the most peaks and generated unique profiles for 29 of the 45 accessions. The remaining 16 fell into eight profile groups of two cultivars each, which were easily resolved with any one of three other primer pairs. For added confidence, strawberry samples at FPS are routinely tested at all six selected markers.
Although the octaploid nature of strawberry complicates interpretation, this system can reliably identify strawberry cultivars at an early developmental stage and distinguish between closely related UC-patented cultivars. This process is used routinely for DNA identity verification of all FPS strawberry mother plants. Furthermore, the SSR technology developed at Foundation Plant Services is available to the public, and private labs are encouraged to use the system.
UC Breeder Selections
Every year, UC strawberry breeders submit to FPS the advanced breeder selections with potential to become new cultivars. These selections are treated using heat and tissue culture techniques to eliminate virus they may have been infected with through exposure to field conditions. The meristem plant for each selection is returned to the breeder for further trials, and a runner from it is held at FPS. This plant is propagated by conventional runner propagation and tested for disease annually. When a breeder decides to release a selection as a new cultivar, FPS staff increases it by conventional runner propagation and moves it to the mother plant greenhouse. By the time a breeder’s selection is released as a new cultivar, it has been at FPS for several years and undergone repeated testing for disease.